Delta™ is the software that empowers our ECZ, ECS, ECA, and ECX series NMR systems. Never has a software package with such powerful control and processing been so easy to use. Multi-dimensional visualization (up to 4D) and processing (up to 8D) are just part of the standard package. Standard Features New!. Object-oriented, multi-dimensional data manipulation. Digital filtering.
Fourier transform. Multi-dimensional phasing. Notch and high-pass filter. Linear prediction, BLIP and FLIP.
Baseline correction. Deconvolution. Automatic peak detection. Data presentation and plotting.
Samsung software upgrade assistant for s6. Data format import and export utilities Delta™ runs on the major platforms: WINDOWS™ and Mac OSX.
Nmr Spectra Services
The Department of Chemistry and Biochemistry has excellent NMR facilities to support its teaching and research programs. A JEOL ECA 500 MHz spectrometer and a JEOL ECS 400 MHz spectrometer are housed at the Patricia Sullivan Science Building in Room 001. The JEOL 500 is built on a 12 T Oxford standard bore magnet and the JEOL 400 on a 9.6 T Jastech magnet.
The JEOL ECA 500 NMR Spectrometer contains a 5 mm tunable PFG probe designed with an outer high frequency coil for 1H and 19F observation and an inner tunable low frequency coil for observation of nuclei including 13C, 15N, 31P, and other esoteric nuclei. The spectrometer contains a Z-axis pulse field gradient system that can operate at 30 G/cm. Gradient shimming makes for convenient operation at high homogeneity. 1-D and 2-D experiments are convenient for both experienced and novice NMR operators using DELTA software with programmed routines including 1-D experiments for a multitude of nuclei as well as COSY, NOESY, HMQC, HMBC, DEPT, HETCOR, HSQC, INEPT, ROESY AND TOCSY. The JEOL ECS 400, our newest spectrometer, has all the capabilities of the JEOL 500 and additionally is equipped with a high sensitivity JEOL Royal probe plus an NMR 24-position sample changer.
The JEOL Delta / NMR support web site provides support for users interested in the Delta NMR Software package and JEOL's NMR Spectrometers. The web site includes Delta NMR software kits for NMR data processing only and kits for JEOL NMR Spectrometer control with data processing. Areas for Delta License Key generation, discussion forums, FAQ's, and documentation downloads are also available on this web site. To get started, simply:.
Free Nmr Software
Registration to this site is moderated - to gain full access, be sure to also submit your spectrometer information so that we can assign you the proper authorization for software downloads and/or license key generation. You will then see content and features available for your account type. If you are a user migrating from ai-sup.jeol.com please. Once your account registration and spectrometer information is reviewed, you will be granted the appropriate access to other areas of the site.
NMR Spectroscopic Data Processing NMR Spectroscopic Data Processing Modern NMR identification uses a range of techniques to provide connectivity and stereochemical information about molecules. In this tutorial, a range of datasets deriving from these techniques are stored on a network file server.
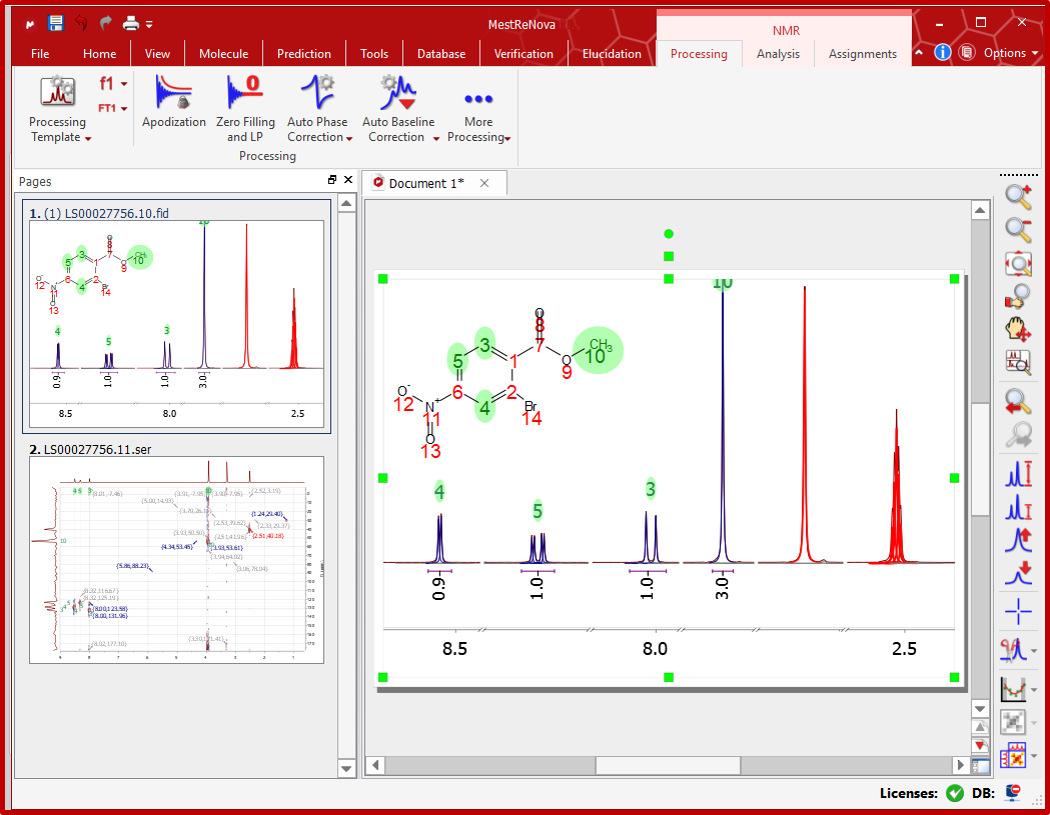
This tutorial illustrates how this data can be processed using appropriate software to provide structural information. Procedure Part I. Processing of Spectrometer Data Data from two principle sources is available for processing, the Bruker spectrometers and the Jeol system. The latter is used for running all Student (1D) NMR samples, the former for more complex (often 2D) data sets. If you are following these instructions to process your laboratory acquired data, use the Jeol option.
If you are following these instructions using the tutorial data for the structure shown below, use the Bruker option. Various programs are available for processing NMR data: Mestrec1D (filename mestrecXY.exe, where XY is the current version number) Mestrec2D (filename mestrecnd.exe) SpecNMR (filename spexnmr.exe) To access these programs, proceed as follows: All the relevant programs are available on the Windows systems located in the Teaching computer room, with shortcuts either as icons on the desktop, or in the start menu. You should also see an NMR Service file server icon, where the data files will be. If you are not using one of these computers, Run Programs/Accessories/Communications/Remote Desktop (Windows 2000/XP only) and specify chas.ch.ic.ac.uk as the remote application server. If the icon to the left is shown, use this shortcut instead. You must remember to log out of these servers when you finish: do not leave yourself logged in, or just quit the remote desktop program without so logging out.
On MacOS X, run the program (found in Applications or the Dock, icon same as above) and specify chas.ch.ic.ac.uk as the remote application servers. You will enter a standard Windows environment, and Mestre-C, SpecNMR and NMR Service icons should be seen on the desktop or from the Start menu. You must remember to log out of these servers when you finish: do not leave yourself logged in, or just quit the remote desktop program without so logging out. Tutorial Data set (Bruker Spectrometers) The Bruker tutorial datasets for the compound shown at the left are located on the local in a folder called NMRData, which contains the following subfolders: 1H is the 400 MHz 1H spectrum 13C is the corresponding 13C data COSY-2D is the 1H/1H correlated 2D data H-C-2D is the 1H/13C correlated 2D data NOE is the 1H NOE experiments The 1D datasets should be opened with the MestreC23 program, the 2D datasets with the MestrecND program (both located on C: Mestrec or D: Mestrec).
Proceed as follows From the 1H sub-folder of NMRData, browse to the file called FID. Note that for this file, you should select a Bruker UXNMR file type. From process, select Fourier transform View/Set limits to view spectrum between predefined values of the chemical shift. Use Tools/Reference and Tools/Integrate, followed by a dragging out of the area of spectrum to be integrated.
Use +/- from the toolbar to change height of spectrum There are numerous other options, which you should explore yourself. The results look akin to this; Repeat for the folder NOE/4-17/pdata/2/1r to see various nOe experiments. The different spectra relate to different pre-irradiations at selected frequencies, and show the difference spectrum resulting. For the 2D plots, proceed as follows; Using MestrecND, go to eg COSY-2D and open the file ser.
Process/Fourier transform/Apodise/SineBell/Apply along t2. You should now see a display comprising a series of vertical stripes. From the menu, change the size to four times its initial value (zero filling eg 256 to 1k) and Apply along t1. The result will be a 2D spectrum. Hit + a few times to increase the size of the contours. Use Tools/Reference F1 and Reference F2 to set the chemical shift to known values, ie TMS.
Proton COSY spectra usually look better if symmetrised (Process menu) and forced to be square. Go to H-C-2D and do the same. In each case the 1H/1H or 1H/13C spectrum is displayed along one frequency axis. The interpretation is that cross peaks in 1H/1H correlation indicate 1H-1H coupling and likewise for 1H/13C. When you have a spectrum to your liking, Copy data to the clipboard and paste into Microsoft Word or other word processor for inclusion in your report.

Do not attempt direct printing, there are no printers provided for the purpose. Laboratory Data sets (Jeol spectrometers) The Jeol datasets derived from samples run as part of laboratory experiments are located in a network drive called 270teaching, subdirectory data on 'chnmr3' in the domain CH1. You may find this folder mounted on the 'Resonance' network drive (270teaching on 'Chnmr3' (R:) The spectra should appear there after the sample has been run.
Note carefully the spectrum code assigned to you. Run the SpecNMR program located in D: SpecNMR (there should be a shortcut on the desktop). Invoke File/Open, select Jeol Delta Format as the file type ( Important!) and click on Drives and browse to R: Data.appropriate sub-folders, Click OK/finish. A list of available files listed in the file dialog box appears. Select the one you want. The screen should shown an FID (Free induction decay).
Select Process/FT+Phase/OK. An (unphased) spectrum appears.
From the Phase floating pane and P0 set, move the slider labelled degrees until the marked peak appears properly phased. Then select P1 and again move the slider until the rest of the spectrum looks properly phased. Finish with Apply. The spectrum can be expanded by dragging out a box for expansion.
Hold the right mouse button down (on a Mac, Control/Shift) and pull out a region on the spectrum to integrate. Repeat as often as necessary. Process/Calibrate to set the reference. Right click to set the point on the spectrum whose chemical shift you know (i.e. TMS) and click OK. If you cannot find a peak whose position you know, enter 8096 (assuming your FID is a 16k one) into the data box and 5.00ppm into the chemical shift box and click OK. The result should be fairly accurate.
The spectrum should look similar to: Edit/copy to clipboard and Paste into Word. Save the Word file either to a removable device (e.g. Pendrive) or to your Network H: drive. Simulation of Spectra (optional) The tutorial relates the following compound, but you should apply the procedure to your own data. Given the structure above, you can simulate the anticipated 1H NMR Spectrum using either:. With the latter, you will have to log into the Application server, and use the built in molecule drawing program, but the operation is fairly self-evident. The results look something like this; Part III.
The Wintorg Tutorials (Optional) From the Windows Start menu, select Wintorg and follow the screen prompts. The program offers you a random selection from a total of 100 unknown compounds.
It is up to you to request the data you feel will most efficiently identify the structure of the compound. What is the future? To find out, go to (C) H S Rzepa, 2002-2003.
Jump to:, Please find complete listing of NMR software. This page is for collecting references on NMR and related software. Information that should be placed here: description, paper reference as well as some publication citing the use of software, download url, set-up instructions etc.
Each piece of software can have it's own separate page and perhaps additional pages linked from within, if necessary. Posts from the original authors are greatly appreciated. Data Acquisition Varian, Inc. Spectrometer software Varian, Inc.
Spectrometer software Bruker BioSpin Gmbh spectrometer software Jeol Ltd. Spectrometer software Data Processing 1D and 2D NMR processing and simulation package. A program written for the old (classic) Macintosh processing, NMR prediction, spectral analysis, structure verification, quantification processing, analysis and simulation for Mac OS X simple processing and display package,.nix and Mac OS X visualisation and processing tool for experimental and simulated (solid-state) NMR data highly flexible toolbox for processing 1D and 2D NMR and EPR spectra under MATLAB, creating high-quality 1D, 2D or 3D plots from the spectra and printing them in every type of format that is supported by MATLAB. Offline Desktop Processing Software that supports all major instrument formats, performs multiplet analysis, and provided structure to spectrum integration for assignment of 1D NMR data.
Offline Desktop Processing Software that supports all major instrument formats, and provides structure to spectrum integration for assignment of 1D and 2D NMR data. NMR Software designed specifically for the synthetic chemists workflow. This software provides quick and easy processing, characterization of multiplets for patents, and journals, as well as a structure verification algorithm that assists the user on the consistency between a proposed chemical structure and an experimental 1H NMR spectrum.
Open source software specialized in DOSY processing. Spectral Analysis Programs NMR Software designed specifically for the synthetic chemists workflow.
This software provides assignment and verification assistance for the 1H NMR spectra of small molecules. Computer- Aided Resonance Assignment runs on Windows, Mac, Linux and Sun operating systems. The Dmfit programs enables fitting of solid state (and liquid) NMR spectra, including 1D and 2D datasets. Edit NMR program: file Viewer for NMR files (Bruker, Ascii.). Quantum mechanical spectral analysis, structure verification, quantification - an open source software package written for the R statistical software. RNMR is designed to simplify the repetitive resonance assignment and quantification tasks associated with metabolomics.
